How Bacteria Repurpose Viruses to Spread Antibiotic Resistance
- Scientists have uncovered a surprising twist in how bacteria share genes—including those that spread antibiotic resistance.
- The findings come from researchers at the John Innes Centre, who studied unusual particles known as gene transfer agents (GTAs).
- These particles act like tiny delivery vehicles, shuttling DNA between cells.
Scientists have uncovered a surprising twist in how bacteria share genes—including those that spread antibiotic resistance. Tiny virus-like particles called gene transfer agents (GTAs), once ancient viral invaders, have been repurposed by bacteria into delivery systems that shuttle DNA between neighboring cells. The study reveals a key control hub of three genes, dubbed LypABC, that triggers bacterial cells to burst open and release these DNA-packed couriers.
The findings come from researchers at the John Innes Centre, who studied unusual particles known as gene transfer agents (GTAs). GTAs resemble bacteriophages (viruses that infect bacteria), but they are no longer harmful invaders. Instead, they are derived from ancient viruses that bacteria have adapted and brought under their own control.
These particles act like tiny delivery vehicles, shuttling DNA between cells. The research shows that when activated by the LypABC gene cluster, bacterial cells undergo a controlled bursting process—essentially exploding—to release GTAs loaded with genetic material. This mechanism enables horizontal gene transfer, a process that can rapidly spread useful traits, including genes that confer resistance to antibiotic drugs used to treat infections.
Understanding Gene Transfer Agents
Gene transfer agents are not typical viruses. While they share structural similarities with bacteriophages, bacteria have domesticated these ancient viral elements over evolutionary time. Rather than causing harm, GTAs now serve as tools for genetic exchange within bacterial communities. This repurposing represents a significant adaptation in microbial biology.

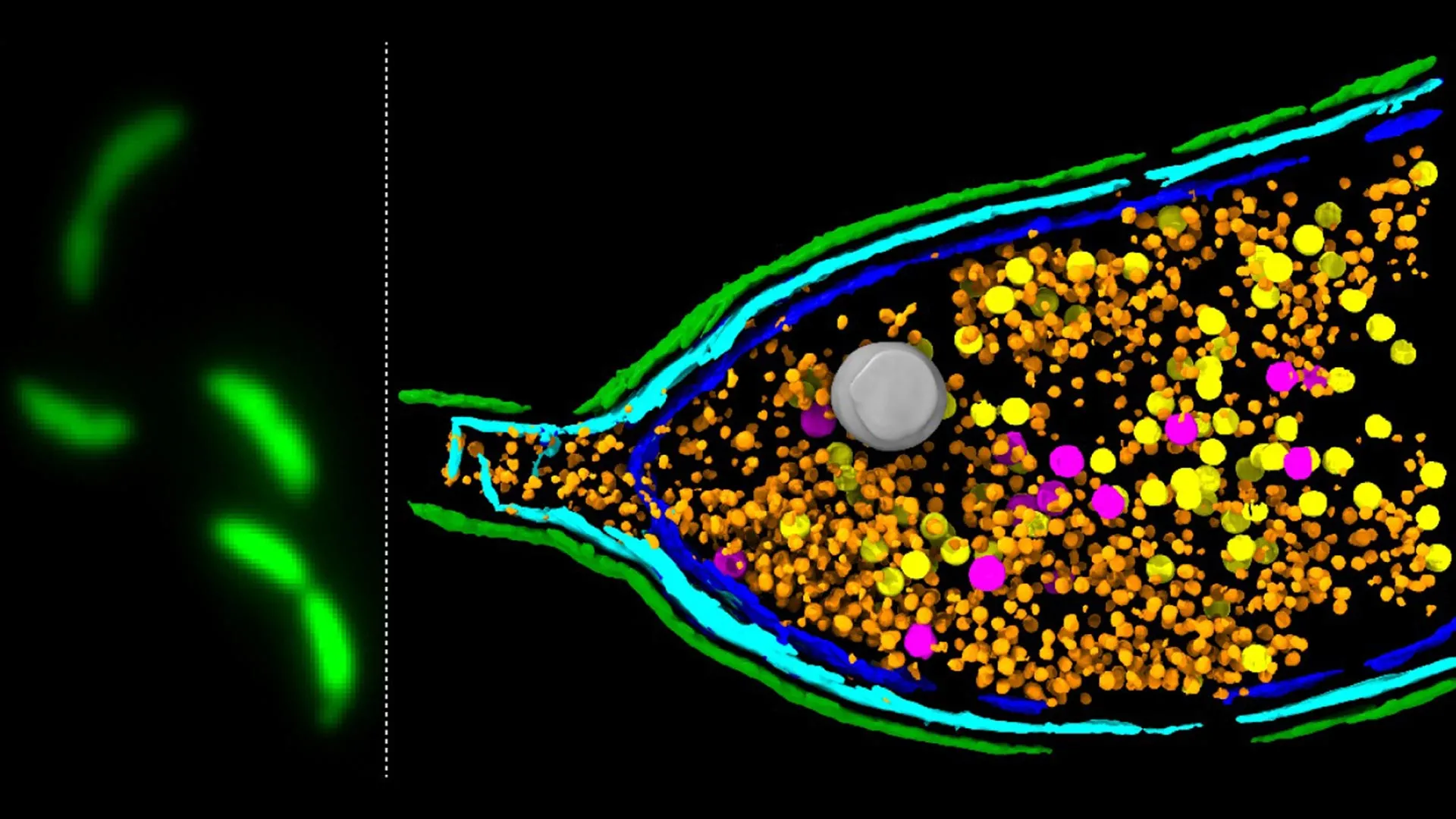
The study used fluorescence microscopy and cryo-electron tomography to observe C. Crescentus bacterial cells engineered to glow green when producing GTAs. These imaging techniques revealed the internal dynamics of GTA production and release, showing how bacterial envelope layers reorganize during the bursting process.
Implications for Antibiotic Resistance
Antimicrobial resistance (AMR) is a growing global health threat. By uncovering how bacteria use GTAs to share resistance genes, scientists gain insight into one mechanism driving the rapid spread of drug-resistant infections. Understanding the LypABC-controlled bursting process may open new avenues for interventions aimed at limiting the spread of resistance without relying solely on new antibiotics.
The research does not suggest immediate clinical applications but highlights a fundamental biological process with relevance to infectious disease dynamics. Further study will be needed to determine whether similar mechanisms operate in pathogenic bacteria and how environmental factors influence GTA-mediated gene transfer.
As antibiotic resistance continues to challenge modern medicine, discoveries like this one provide essential pieces of the puzzle. By revealing how bacteria hijack their own systems to share DNA, scientists move closer to understanding the full scope of microbial adaptation and evolution in the face of antimicrobial pressure.
