Predicting Bacterial Antiphage Systems Using Machine Learning
- Researchers have utilized machine learning and deep learning to identify thousands of previously unknown bacterial immune defense systems, revealing a level of molecular diversity in bacterial immunity that...
- The findings, detailed in research published in Science on April 2, 2026, demonstrate that bacteria possess a vast array of antiphage systems designed to protect them from viral...
- One group of researchers, led by DeWeirdt et al., developed a specific model known as DefensePredictor to discover these systems.
Researchers have utilized machine learning and deep learning to identify thousands of previously unknown bacterial immune defense systems, revealing a level of molecular diversity in bacterial immunity that remains largely uncharacterized.
The findings, detailed in research published in Science on April 2, 2026, demonstrate that bacteria possess a vast array of antiphage systems designed to protect them from viral infections. To uncover the extent of this diversity, two independent research groups developed complementary models that leverage genomic context and protein sequences to predict these defense systems at scale.
Machine Learning Approaches to Bacterial Immunity
One group of researchers, led by DeWeirdt et al., developed a specific model known as DefensePredictor
to discover these systems. In a separate but parallel effort, Ernest Mordret et al. Also developed machine learning models to predict antiphage defense systems across large datasets.
These researchers employed protein and genomic language models to systematically predict antiphage function. By analyzing the bacterial pangenome, the models were able to achieve a precision rate of up to 99% and a recall rate of 92%.
Scale and Scope of the Discovery
The application of these models to more than 32,000 bacterial genomes has significantly expanded the known repertoire of antiphage systems. The data indicates that approximately 1.5% of bacterial genomes are devoted to these defense mechanisms.

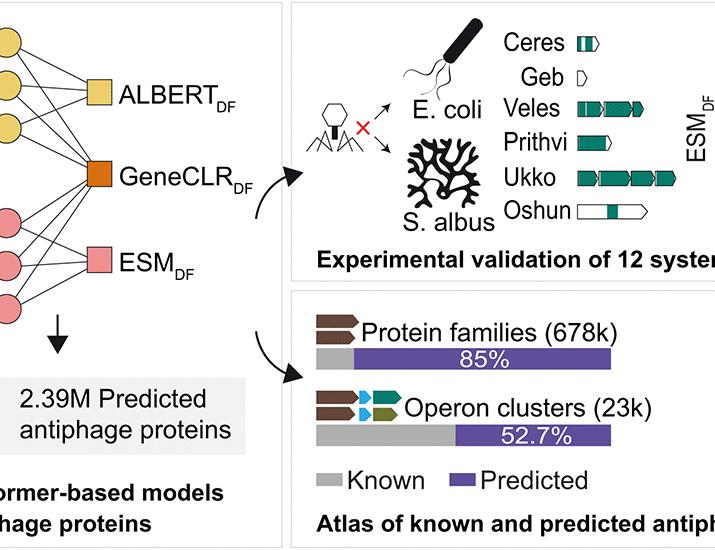
Despite the success of the predictive models, much of the discovered diversity is still not understood. More than 85% of the predicted protein families involved in these defense systems remain uncharacterized.
To facilitate further study, the researchers have provided an interactive catalog containing more than 19,000 candidate operon families. This catalog is intended to serve as a resource for future experimental follow-up to determine the exact functions of these systems.
Experimental Validation
To verify the accuracy of the machine learning predictions, the researchers conducted experimental validations using Escherichia and Streptomyces. These experiments led to the actual discovery of 12 antiphage systems, confirming that the models can successfully identify functional defense mechanisms in living bacteria.
The research suggests that the majority of the molecular diversity used by bacteria to fight off viruses has not yet been described by science, providing a new foundation for the systematic exploration of bacterial immunity.
Potential Implications for Genetic Editing
The discovery of these unknown immune systems may have applications beyond basic microbiology. Insights into how bacteria defend themselves against phages could potentially impact the field of genetic editing.
By understanding the diverse mechanisms bacteria use to recognize and neutralize foreign genetic material, scientists may find new tools or methods to improve the precision and efficiency of genome editing technologies.
- The study used complementary machine learning and deep learning models to predict antiphage functions.
- Over 32,000 bacterial genomes were analyzed, showing that 1.5% of these genomes are dedicated to defense.
- The researchers identified an interactive catalog of over 19,000 candidate operon families.
- Experimental validation in Escherichia and Streptomyces confirmed the existence of 12 new antiphage systems.
- More than 85% of the protein families predicted by the models are currently uncharacterized.
